North Carolina Macular Dystrophy
All content on Eyewiki is protected by copyright law and the Terms of Service. This content may not be reproduced, copied, or put into any artificial intelligence program, including large language and generative AI models, without permission from the Academy.
Disease
North Carolina Macular Dystrophy (NCMD)
Alias: Lefler, Wadsworth and Sidbury Syndrome, Hereditary Macular Degeneration and Amino-aciduria, Dominant Progressive Foveal Dystrophy, Central Areolar Pigment Epithelial Dystrophy (CAPED), Autosomal Dominant Central Pigment Epithelial and Choroidal Degeneration
Etiology
Inherited, autosomal dominant

Symptoms
About a third of the patients are asymptomatic. A third have mild central visual impairment. Another third have mild to moderate central vision impairment. A few have severe central vision impairment typically due to choroidal neovascular membrane (CNVM) formation.
History
NCMD is an autosomal dominant, congenital, completely penetrant, non-progressive macular malformation first reported 50 years ago in a large family in North Carolina. This family’s retinal findings were initially described by Lefler, Wadsworth and Sidbury as “hereditary macular degeneration and amino-aciduria” and subsequently referred to as the Lefler Wadsworth Sidbury Syndrome.[2][3] The “aminoaciduria” component, was later found to be inconsistent making the “syndrome” a misnomer.[3] This same family was reported in a cross-sectional study 3 years later as “a new dominant progressive foveal dystrophy.” [3]
Twenty years later, Small encountered this same family and reevaluated and defined the disease phenotype in many of the same individuals. Only one family member lost vision over time, which appeared to be due to the development of CNVM. This disease and this family were noted in Gass’ original Atlas of Macula Diseases as “North Carolina Macular Dystrophy” for its founder effect.[4] This has been the popular name ever since but is also inaccurate and yet another misnomer associated with this disease.[5] [6] [7] Other macular dystrophies such as “Central Areolar Pigment Epithelial Dystrophy (CAPED)” and “Autosomal Dominant Central Pigment Epithelial and Choroidal Degeneration (ADCPECD)” as described by Leveille et al. were subsequently shown by Small et al. to actually be genealogical branches of the original NCMD family.[7][8] Indeed, Leveille et al. used the presence of peripheral drusen to split ADCPECD from Lefler’s NCMD.[8] Therefore, “lumping” disease phenotypes together has proven to be more accurate than “splitting.”[9][10] Although considered rare, NCMD has been reported worldwide in over 50 families.[5] [6] [7][11] [12] [13] [14] [15] [16] [17] [18] [19] [20] [21] [22] [23] [24] [25] [26] [27] [28] [29] [30] [31] [32] [33] [34] [35] [36] [37][38] Affected families have been reported in the United States, Europe, Central America, Australia, New Zealand, Korea, Turkey and China.[11] [12] [13] [14] [15] [16] [17] [18] [19] [20] [21] [22] [23] [24] [25] [26] [27] [28] [29] [30] [31] [32] [33] [34] [35] [36] [37] Hence, “North Carolina macular dystrophy” is a gross misnomer.
Pathophysiology
NCMD is a congenital developmental abnormality of the macula. Most affected individuals carry a mutation in noncoding DNA found on the MCDR1 locus of Chromosome 6. The implicated variants alter a DNase1 hypersensitivity binding site upstream of the PRDM13 gene, which is proposed to dysregulate the PRDM13 gene through altered chromatin assembly. [39] The PRDM13 gene product is a transcription factor expressed in developing neural retina, particularly in amacrine cells.[40] The MCDR3 locus on chromosome 5 has also separately been implicated in causing NCMD. At this locus in diseased individuals, Cirpriani et al have reported discovery of a duplicated region containing the IRX1 gene, whose product is also a developmental transcription factor not exclusive to the nervous system. [39] Although detailed mechanisms of macular degeneration as a result of these mutations remain unclear, one theory is that these mutations alter chromatin folding causing dysfunctional structural variants.
Diagnosis
Clinical Diagnosis
There is a great deal of variable expressivity in the appearance of affected maculae. In general, the macular lesions are congenital and bilaterally symmetrical. The appearance of the lesions range from central small/intermediate drusen (grade 1), to confluent drusen, to central vitelliform lesions (grade 2), to large central defects with atrophy that bow back into the choroid and sclera (grade 3), often with surrounding subretinal fibrosis (most likely from CNVM). Figure 1 demonstrates the appearance of typical macular lesions from an affected individual over three decades of follow up. [41]
Visual acuity is generally better than would be expected given the appearance of the lesions but has been reported to range from 20/20 to worse than 20/400. The visual acuity generally stabilizes throughout the patient’s lifetime unless the patient develops a CNVM in the area of fixation (typically slightly nasal to the fovea). Color vision is typically normal, which aids in differentiating NCMD from other inherited retinal disorders.
Other Exam
Because the macular lesions (grade 3 colobomatous-like) are typically centered slightly temporal to the macula, some patients may have the appearance of strabismus as they adjust fixation. Some affected families have shown “claw hand” malformations or hearing loss. Examining other family members can reveal the variable expressivity of involved mutations and can greatly facilitate diagnosis.
Molecular Diagnosis
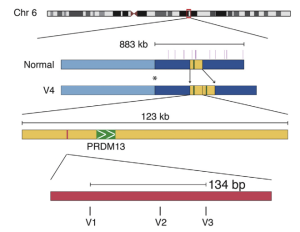
Known causative mutations of NCMD have been discovered in the MCDR1 locus on chromosome 6 (Figure 2) and the MCDR3 locus on chromosome 5. Clinical diagnosis can be confirmed with DNA sequencing for known mutations (see table below). It must be emphasized that failing to detect a known mutation does not exclude a diagnosis of NCMD. Of note, commercial labs offering genetic testing for inherited retinal disorders currently only detect mutations on the MCDR1 chromosome 6 locus and not on the MCDR3 chromosome 5 locus, which are more prevalent in the European population. Clinical judgment should determine whether further specialized genetic testing, such as whole genome or exome sequencing should be pursued.
| Variant Number | phenotype | Type of Variant | Chromosomal Position (Hg19) | Nucleotide Change | Geographic location | Found by: | PCR sequencing primer sequences | ||
| MCDR1 | Chromosome 6 | ||||||||
| V1 | NCMD / MCDR1 | SNP | 6: 100040906 | c.-14005G>T | G>T | USA | Small et al 2017 | F: 5'-GCATTCCCTAAAGCACTTGACC-3'
R: 5'-GATAGCTACCCCTCCTCTGAATG-3' | |
| V2 | NCMD / MCDR1 | SNP | 6: 100040987 | c.-13924G>T | G>C | France/ Germany/ USA /UK | Small et al 2017 | F: 5'-CTGATCATTTGAATCAAGGCAG-3'
R: 5'-CAGCACTTGCACATTTGTGTC-3' | |
| V3 | NCMD / MCDR1 | SNP | 6: 100041040 | c.-13871C>T | C>T | China | Small et al 2017 | F: 5'-CACTGGAAAAATTATGTGGAAATC-3'
R: 5'-GAGTAATTAATGAAGTTGACAAGTTG-3' | |
| V4 | NCMD / MCDR1 | Tandem DUP | 6: 100020205-100143306 | 123,101 bp DUP | Belize | Small et al 2017 | F: 5'-GATAAATCATATCTTAGACCGC-3'
R: 5'-CTCATGCCTATAATCCCAGCAC-3' | ||
| V6 | NCMD / MCDR1 | Tandem DUP | 6: 99996226-100065137 | 69,912 bp DUP | USA | Brown et al 2019 | F: 5'-TGTAAAACGACGGCCAGTCTGGCTCAGTACCAACTCCTG-3'
R: 5'-CAGGAAACAGCTATGACCGGAACAGAATACTGGGTCCTTT-3' F: 5'-TGTAAAACGACGGCCAGTCATGGGAAACATTTTATCAGC-3' R: 5'-CAGGAAACAGCTATGACCTGTTGATGTTTCTTGGCCTCC-3' | ||
| V7 | NCMD / MCDR1 | Tandem DUP | 6: 99984309-100082698 | 98,389 bp DUP | France / Italy | Manes et al 2019 / Small et al | F: 5'-AGGCACCTGAGGTAAGCAGA-3'
R: 5'-CCTGACCTCAGGTGATCTGC-3' | ||
| V10 | PBRCA | SNP | 6:100046804 | c.-8107T>C NM_021620.3 | T>C | UK | Silva et al 2019 | ||
| V11 | severe NCMD / PBRCA- | SNP | 6: 100046783 | c.-8128A>C NM_021620.3 | A>C | Asian / Egypt | Silva et al 2019 / Small et al | ||
| V12 | NCMD / MCDR1 | Tandem DUP | 6: 99112389-99168616 | 56,228 bp DUP | Turkey | Small et al 2020 /Vance/Ozge/deBaere | |||
| V13 | NCMD / MCDR1 | SNP CHF? | 6: 100040974 | A>C | Namburi et al 2020 | ||||
| MCDR3 | Chromosome 5 | ||||||||
| V5 | NCMD / MCDR3 | Tandem DUP | 5: 3587901-4486027 | 898,126 bp DUP | Denmark | Small et al 2017 | F: 5'-GTTTTCACGAAAGTGCAAAGG-3'
R: 5'-GGGGTGGAAGAGAAGAGAGG-3' | ||
| V8 | NCMD / MCDR3 | Tandem DUP | 5:4391377–4436535 | 45,158 bp DUP | UK / German | Cipriani et al 2019 | F:5′-TTTGCTTGATCAATTCTGCTG-3′
R:5′-TTCTCAGTTGGAAGAGCACAAA-3′ | ||
| V9 | NCMD / MCDR3 | Tandem DUP | 5:4396927–4440442 | 43,515 bp DUP | UK / German | Cipriani et al 2019 | F: 5′-TTGTGGACTGAGCAAGCAAG-3
R:5′-GGAGCAGAAGTTAAATGTGGAGA-3′ | ||
Diagnostic procedures
Fundus exam, fundus photography, fluorescein angiography, indocyanine green (ICG) angiography, all show findings consistent with the findings noted in “DIAGNOSIS”. Additionally an active CNVM may be found.
Electroretinography (ERG): Full field ERG is typically normal. This test is useful in distinguishing NCMD from other IRDs. However, there is one report by Manes et al[36] of an abnormality with the oscillatory potentials (OPs) electrooculography consistent with amacrine cell dysfunction. However, “normative data” of OPs is lacking in the literature and what constitutes an abnormal OP recording is not well-established.
Electrooculogram (EOG) recordings, are typically normal although Small et al has reported several patients with an abnormally reduced Arden ratio with a normal ERG. The traditional teaching is that a normal ERG with an abnormal EOG suggests the diagnosis of Best Vitelliform Macular Dystrophy (BVMD). This classic teaching does not account for the same finding in some patients with NCMD.
Differential diagnosis
- Congenital toxoplasmosis
- Best vitelliform macular dystrophy (BVMD)[42]
- Age-related macular degeneration (AMD)
- Torpedo maculopathy
- Cone dystrophy (CORD5 CORD6)
- Progressive Bifocal RetinoChoroidal Degeneration (PBRCA)
General treatment
Periodic monitoring for the development of CNVMs. Studies have demonstrated that anti-VEGF injections work well to address complications from CNVM in NCMD.[43] Refractive correction, low vision aids, and cataract surgery can help patients depending on their ocular status.
Medical therapy
Anti-VEGF injections to manage CNVM in affected patients.
Surgery
Prior to the development of intravitreal anti-VEGF injections, submacular surgery was rarely performed. For patients with macular lesions affecting fixation with resultant appearance of strabismus, strabismus surgery has not been reported to have much success in improving visual function or appearance.
Complications
The development of CNVMs can cause progressive vision loss.
Prognosis
Prognosis is generally good for maintaining useful and stable vision throughout the life of the patient. Even patients with severe appearing grade 3 colobomatous-like lesions can maintain 20/40 visual acuity.
References
- ↑ Jump up to: 1.0 1.1 Small KW, DeLuca AP, Whitmore SS, Rosenberg T, Silva-Garcia R, Udar N, Puech B, Garcia CA, Rice TA, Fishman GA, Héon E, Folk JC, Streb LM, Haas CM, Wiley LA, Scheetz TE, Fingert JH, Mullins RF, Tucker BA, Stone EM. North Carolina Macular Dystrophy Is Caused by Dysregulation of the Retinal Transcription Factor PRDM13. Ophthalmology 123:9-18, 2016
- ↑ Lefler WH, Wadsworth JA, Sidbury JB. Hereditary macular degeneration and amino-aciduria. Am J Ophthalmol. 1971;71:224–230.
- ↑ Jump up to: 3.0 3.1 3.2 Frank HR, Landers MB, Williams RJ, Sidbury JB. A new dominant progressive foveal dystrophy. Am J Ophthalmol. 1974;78:903–916.
- ↑ Agarwal A. Gass’ Atlas Of Macular Diseases. 5th ed. Edinburgh, Scotland: Elsevier Saunders; 2012.
- ↑ Jump up to: 5.0 5.1 Small KW. North Carolina macular dystrophy: revisited. Ophthalmology 96:1989,1747-1754.
- ↑ Jump up to: 6.0 6.1 Small KW, Killian J, McLean W. North Carolina's dominant progressive foveal dystrophy. How progressive is it? British Journal of Ophthalmology 75:1991,401-406
- ↑ Jump up to: 7.0 7.1 7.2 Small KW, Hermsen V, Gurney N, Fetkenhour CL, Bresnick G, Folk JC: North Carolina macular dystrophy (NCMD) and central areolar pigment epithelial dystrophy (CAPED), one family, one disease. Archives of Ophthalmology 110:1991,515-518.
- ↑ Jump up to: 8.0 8.1 Leveille AS, Morse PH, Kiernan JP. Autosomal dominant central pigment epithelial and choroidal degeneration. Ophthalmology. 1982 Dec;89(12):1407-13. doi: 10.1016/s0161-6420(82)34621-7. PMID: 7162784.
- ↑ Traboulsi EI. To lump or to split? Ophthalmic Pediatric Genetics. 1993 Dec;14(4):141-2. doi: 10.3109/13816819309042912.
- ↑ Freund KB, Sarraf D, Mieler WF, Yannuzzi LA, Shields CL. The Retinal Atlas. Philadelphia, PA: Elsevier:2017.
- ↑ Jump up to: 11.0 11.1 Rabb M, Small KW, Mullen L, Yelchits L, Udar, N. North Carolina macular dystrophy maps to the MCDR1 locus (MCDR1) phenotype in Central America. Am J Ophthalmol 125:502-508, 1998.
- ↑ Jump up to: 12.0 12.1 Small KW, Puech B, Mullen L, Yelchits L. North Carolina macular dystrophy phenotype in France maps to the MCDR1 locus. Molecular Vision 3:1, 1997.
- ↑ Jump up to: 13.0 13.1 Small KW. North Carolina macular dystrophy. Transactions of the American Ophthalmological Society XCVI: 926-961, 1998.
- ↑ Jump up to: 14.0 14.1 Voo I, Glasgow B, Flannery J, Udar U, Small KW. Clinicopathologic correlation of North Carolina macular dystrophy. Am J Ophthalmol. 2001;132(6):933-5.
- ↑ Jump up to: 15.0 15.1 Small KW, Voo I, Glasgow B, Flannery J, Udar N. Clinicopathologic correlation of North Carolina macular dystrophy. Trans Am Oph Soc 99: 233-238, 2001
- ↑ Jump up to: 16.0 16.1 Kiernan DF, Shah RS, Hariprasad SM, Grassi MA, Small KW, Kiernan JP, Mieler WF. Thirty Year Follow-up of an African-American Family with North Carolina Macular Dystrophy (MCDR1). Ophthalmology 2011;118:1435–1443.
- ↑ Jump up to: 17.0 17.1 Small KW, Puech B, Mullen L, Yelchits S. North Carolina macular dystrophy phenotype in France maps to the MCDR1 locus. Mol Vis. 1997;3:1.
- ↑ Jump up to: 18.0 18.1 Small KW, Garcia CA, Gallardo G, et al. North Carolina macular dystrophy (MCDR1) in Texas. Retina. 1998;18:448e452.
- ↑ Jump up to: 19.0 19.1 Rabb MF, Mullen L, Yelchits S, et al. A North Carolina macular dystrophy phenotype in a Belizean family maps to the MCDR1 locus. Am J Ophthalmol. 1998;125:502e508.
- ↑ Jump up to: 20.0 20.1 Small K, Small L, Tran E, Rao R, Shaya F. Multimodal Imaging and Functional Testing in a North Carolina Macular Disease Family: Toxoplasmosis, Fovea Plana, and Torpedo Maculopathy Are Phenocopies. Ophthalmology Retina, Volume 3, Issue 7, 607-614.
- ↑ Jump up to: 21.0 21.1 Small KW, Vincent A, Knapper CL, Shaya F. Congenital toxoplasmosis as one phenocopy of North Carolina Macular Dystrophy (NCMD/MCDR1). American Journal of Ophthalmology Case Reports, Volume 15, 2019, 100521. https://doi.org/10.1016/j.ajoc.2019.100521.
- ↑ Jump up to: 22.0 22.1 Small KW, Agemy S, Shaya FS. Terminology of MCDR1: What’s in a name? JAMA Ophthalmol. 2016;134:355e356.
- ↑ Jump up to: 23.0 23.1 Benjamin Bakall, MD, PhD, J. Shepard Bryan III, MD, Edwin M. Stone, MD, PhD, Kent W. Small, MD. Choroidal neovascularization in North Carolina macular dystrophy responsive to anti–vascular endothelial growth factor therapy. Retin Cases Brief Rep. 2018 Oct 31. doi: 10.1097/ICB.0000000000000838. PubMed PMID: 30383557
- ↑ Jump up to: 24.0 24.1 Small KW, Weber JL, Roses AD, Pericak-Vance M. North Carolina macular dystrophy maps to chromosome 6. Genomics 13:1992,681-685.
- ↑ Jump up to: 25.0 25.1 Small KW, Weber J, Pericak-Vance MA. MCDR1 (North Carolina macular dystrophy) map to 6q14-q16. Ophthalmic Pediatrics and Genetics 14:143-150, 1993.
- ↑ Jump up to: 26.0 26.1 Small KW, Weber JL, Pericak-Vance MA, Vance J, Hung W, Roses AD. Exclusion map of North Carolina macular dystrophy using RFLPs and microsatellites. Genomics 11:1991,763-766.
- ↑ Jump up to: 27.0 27.1 Small KW, Udar N, Yelchits S, et al. North Carolina macular dystrophy (MCDR1) locus: a fine resolution genetic map and haplotype analysis. Mol Vis. 1999.
- ↑ Jump up to: 28.0 28.1 Small KW, DeLuca AP, Whitmore SS, et al. North Carolina macular dystrophy is caused by dysregulation of the retinal transcription factor PRDM13. Ophthalmology. 2016;123:9e18.
- ↑ Jump up to: 29.0 29.1 Pauleikhoff D, Sauer CG, Müller CR, et al. Clinical and genetic evidence for autosomal dominant North Carolina macular dystrophy in a German family. Am J Ophthalmol.1997;124:412e415.
- ↑ Jump up to: 30.0 30.1 Rohrschneider K, Blankenagel A, Kruse FE, et al. Macular function testing in a German pedigree with North Carolina macular dystrophy. Retina. 1998;18:453e459.
- ↑ Jump up to: 31.0 31.1 Reichel MB, Kelsell RE, Fan J, et al. Phenotype of a British North Carolina macular dystrophy family linked to chromosome 6q. Br J Ophthalmol. 1998;82:1162e1168.
- ↑ Jump up to: 32.0 32.1 Rosenberg T, Roos B, Johnsen T, et al. Clinical and genetic characterization of a Danish family with North Carolina macular dystrophy. Mol Vis. 2010;16:2659e2668.
- ↑ Jump up to: 33.0 33.1 Kim SJ, Woo SJ, Yu HG. A Korean family with an early-onset autosomal dominant macular dystrophy resembling North Carolina macular dystrophy. Korean J Ophthalmol. 2006;20: 220e224.
- ↑ Jump up to: 34.0 34.1 Small KW, Van de Sompele S, Nuytemans K, et al. A novel duplication involving PRDM13 in a Turkish family supports its role in North Carolina macular dystrophy (NCMD/MCDR1). Mol Vis. 2021;27:518-527.
- ↑ Jump up to: 35.0 35.1 Seiple W, Szlyk JP, Paliga J, Rabb MF. Perifoveal Function in Patients with North Carolina Macular Dystrophy: The Importance of Accounting for Fixation Locus. Invest. Ophthalmol. Vis. Sci. 2006;47(4):1703-1709. doi: 10.1167/iovs.05-0659.
- ↑ Jump up to: 36.0 36.1 36.2 Manes G, Joly T, Smirnov S, et al. A novel duplication of PRMD13 causes North Carolina macular dystrophy: overexpression of PRDM13 orthologue in Drosophila eye reproduces the human phenotype. Hum Mol Genet. 2017;26:4367e4374.
- ↑ Jump up to: 37.0 37.1 Bowne SJ, Sullivan LS, Wheaton DK, et al. North Carolina macular dystrophy (MCDR1) caused by a novel tandem duplication of the PRDM13 gene. Mol Vis. 2016;22:1239e1247.
- ↑ Green DJ, Lenassi E, Manning CS, McGaughey D, Sharma V, Black GC, Ellingford JM, Sergouniotis PI. North Carolina Macular Dystrophy: Phenotypic Variability and Computational Analysis of Disease-Associated Noncoding Variants. Invest Ophthalmol Vis Sci. 2021 Jun 1;62(7):16. doi: 10.1167/iovs.62.7.16.
- ↑ Jump up to: 39.0 39.1 Cipriani V, Silva RS, Arno G, et al. Duplication events downstream of IRX1 cause North Carolina macular dystrophy at the MCDR3 locus. Sci Rep. 2017;7(1):7512. Published 2017 Aug 8. doi:10.1038/s41598-017-06387-6.
- ↑ Watanabe S, Sanuki R, Sugita Y, Imai W, Yamazaki R, Kozuka T, Ohsuga M, Furukawa T. Prdm13 regulates subtype specification of retinal amacrine interneurons and modulates visual sensitivity. J Neurosci. 2015 May 20;35(20):8004-20. doi: 10.1523/JNEUROSCI.0089-15.2015. PMID: 25995483; PMCID: PMC6795186.
- ↑ Small KW, DeLuca AP, Whitmore SS, Rosenberg T, Silva-Garcia R, Udar N, Puech B, Garcia CA, Rice TA, Fishman GA, Héon E, Folk JC, Streb LM, Haas CM, Wiley LA, Scheetz TE, Fingert JH, Mullins RF, Tucker BA, Stone EM. North Carolina Macular Dystrophy Is Caused by Dysregulation of the Retinal Transcription Factor PRDM13. Ophthalmology 123:9-18, 2016
- ↑ Small KW, Jampol LM, Bakall B, Small L, Wiggins R, Agemy S, Udar N, Avetisjan J, Vincent A, Shaya FS. Best Vitelliform Macular Dystrophy (BVMD) is a phenocopy of North Carolina Macular Dystrophy (NCMD/MCDR1). Ophthalmic Genet. 2021 Dec 13:1-11. doi: 10.1080/13816810.2021.2010771. Epub ahead of print. PMID: 34895015.
- ↑ Bakall B, Bryan JS 3rd, Stone EM, Small KW. CHOROIDAL NEOVASCULARIZATION IN NORTH CAROLINA MACULAR DYSTROPHY RESPONSIVE TO ANTI-VASCULAR ENDOTHELIAL GROWTH FACTOR THERAPY. Retin Cases Brief Rep. 2021 Sep 1;15(5):509-513. doi: 10.1097/ICB.0000000000000838. PMID: 30383557.

